Professional Manufacturer of Biomagnetic Beads

A Key Guide to Maximizing the Efficiency of DNA Fragment Selection
Foreword
Possessing high-performance DNA fragment selection magnetic beads lays a solid foundation for experimental success. However, the final outcome of a selection experiment is the combined result of the beads’ inherent properties and the operator’s precise technique.
Even the most advanced beads can lead to selection failure if used with improper protocols. Mastering the following six key operational considerations is the practical key to ensuring you achieve high-resolution, high-recovery, and highly reproducible selection results.
Consideration 1:
Precise Preparation and Verification of Key Buffers – Controlling the “Chemical Code” of Separation
Operational Point: The PEG (polyethylene glycol) concentration, type and concentration of salt ions, and pH in the binding and elution buffers are the core variables controlling the selection window. Use high-purity reagents, prepare solutions via accurate weighing and pH meter calibration, and aliquot into small volumes for storage to avoid concentration changes from repeated freeze-thaw cycles or long-term storage.
Scientific Basis: PEG concentration affects the steric exclusion effect, while salt concentration governs the strength of the “salt-bridge” binding. Together, they determine the binding affinity threshold between DNA of different lengths and the beads. Minor deviations (e.g., a 1% change in PEG concentration) can shift the selection window center by 10-20bp. Therefore, the accuracy of buffer preparation is the first critical checkpoint for selection success or failure.
Consideration 2:
Strict Optimization of Bead-to-Sample Ratio – Avoiding “Overloading” and “Waste”
Operational Point: Do not use a fixed bead addition ratio for all samples. Based on the target fragment size range, the total amount of input DNA, and its length distribution, perform optimization experiments within the recommended range in the protocol to determine the optimal bead volume (typically expressed as a volume ratio).
Scientific Basis: The binding sites on the beads are limited. Insufficient bead volume leads to saturation, causing leftover target DNA to be discarded and lowering recovery. Excessive bead volume may cause non-specific adsorption of short-fragment impurities that should be removed, reducing selection purity and resolution. Optimizing the ratio finds the equilibrium point for maximizing both yield and purity.
Consideration 3:
Uniformity of Temperature and Time Control – Ensuring Reaction Consistency
Operational Point: Perform the binding and elution incubation steps on a thermostatic mixer to ensure all sample tubes are at a completely consistent and constant temperature (typically room temperature, 25°C) and receive uniform mixing. Strictly and precisely time each incubation step.
Scientific Basis: The binding and dissociation of DNA to/from the beads are thermodynamically and kinetically controlled processes. Temperature fluctuations alter the reaction equilibrium constant, uneven mixing affects mass transfer efficiency, and timing differences cause variations in reaction progression. Any inconsistency directly translates into intra-batch or inter-batch variability in selection results, compromising experimental reproducibility.
Consideration 4:
Gentle Handling to Avoid Physical Shearing – Protecting Target Molecule Integrity
Operational Point: When handling samples containing DNA-bead complexes or long DNA fragments, always use wide-bore pipette tips and handle gently. Avoid vigorous vortexing, high-speed shaking, or repeated forceful pipetting.
Scientific Basis: High-molecular-weight DNA strands are extremely sensitive to fluid shear forces. Aggressive physical manipulation can cause non-specific shearing of the target long DNA fragments. This not only reduces the recovery of the target fragments but also generates unintended short-fragment contaminants, severely compromising the homogeneity and purity of the selected product.
Consideration 5:
“Fractionated Collection” Strategy for the Elution Step – Refined Acquisition of Target Product
Operational Point: When eluting the target DNA, it is recommended to adopt a strategy of adding the elution buffer (e.g., nuclease-free water or TE buffer) in small, multiple aliquots (e.g., twice), collecting the eluate from each step separately.
Scientific Basis: The first elution typically contains the core target fragments with the most optimal binding affinity, offering the highest purity. The second elution may contain fragments that bound slightly more tightly, which are often slightly larger than the target range. After fractionated collection, concentration measurement and fragment distribution analysis (e.g., by microfluidic capillary electrophoresis) allow for selective pooling of the desired fractions, yielding a narrower and more precise selected product than a single bulk elution.
Consideration 6:
Implementing Quality Control Checkpoints Throughout the Process – Replacing Judgment with Data
Operational Point: For every formal selection experiment, parallel quality control of both the input DNA before selection (Input) and the recovered product after selection (Output) is mandatory. Relying solely on spectrophotometer readings for assessing success is a significant misconception.
Scientific Basis: Using a microfluidic electrophoresis system to analyze pre- and post-selection samples is the “gold standard” for objectively evaluating selection performance. It visually reveals: 1) Efficiency: Recovery status of the target peak; 2) Resolution: Whether the product peak is sharp and tightly distributed; 3) Specificity: Whether impurities like primer dimers have been effectively removed.
Product Introduction
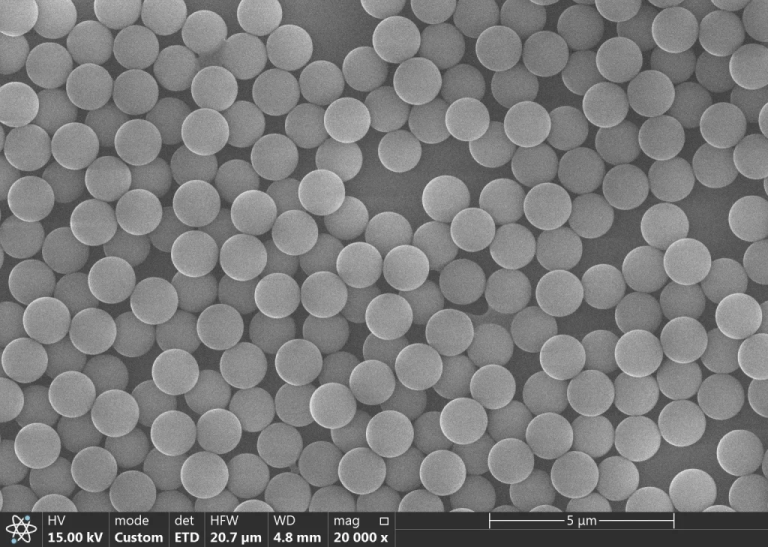
Leveraging mature bead design and manufacturing processes, Shanghai Lingjun Biotechnology Co., Ltd. has developed its high-performance FR0015 Series Silica Magnetic Beads. With an average particle size of 300nm, strong magnetic properties, excellent monodispersity, and a half-sedimentation time exceeding 1 hour, this product line is primarily suited for:
Serum/Plasma cell-free DNA/RNA fragment extraction
DNA fragment sorting/selection
Applications requiring excellent bead suspension stability and co-extraction of DNA/RNA fragments across a broad size range.
Conclusion
Exceptional experimental results stem from rigorous control over every detail. Combining superior tools with standardized operation is the way to unleash the full potential of silica-based magnetic beads for selection.
We are committed not only to providing fragment selection beads that meet the most stringent performance requirements but also to being your reliable partner in experimentation. We offer detailed, standardized operational protocols and robust technical support to help you avoid common pitfalls and consistently produce high-quality selection results. Let’s work together to ensure every step, from sample to data, is clear, precise, and trustworthy, with scientific rig
Supplier
Shanghai Lingjun Biotechnology Co., Ltd. was established in 2016 which is a professional manufacturer of biomagnetic materials and nucleic acid extraction reagents.
We have rich experience in nucleic acid extraction and purification, protein purification, cell separation, chemiluminescence, and other technical fields.
Our products are widely used in many fields, such as medical testing, genetic testing, university research, genetic breeding, and so on. We not only provide products but also can undertake OEM, ODM, and other needs. If you have a related need, please feel free to contact us .